Peaks Code
The "peaks" code for attempting to go from a map to a structure is one of the most complicated parts of the edm package, and the one where a substantial amount of user input is needed and there is no single correct approach. What you are trying to do is reduce a map with peaks at the probable location of atoms or atomic columns to a reasonable structure model. The code knows very little about the specific problem that you have, for instance what constraints there are on the chemistry and/or possible structures from other information that you have. Your mission, should you agree to accept it, is to use the code to try and find a structure consistent with all the other information available. An example exercise can be found by following this link:
Peaks is a rather
sophisticated program, and requires the attention of the user in order to
perform a successful structure completion. You must never abandon your chemical
intuition. Pay attention to bond distances, charge balance constraints, etc.
The main window, shown below, has a large number of different controls to help you.
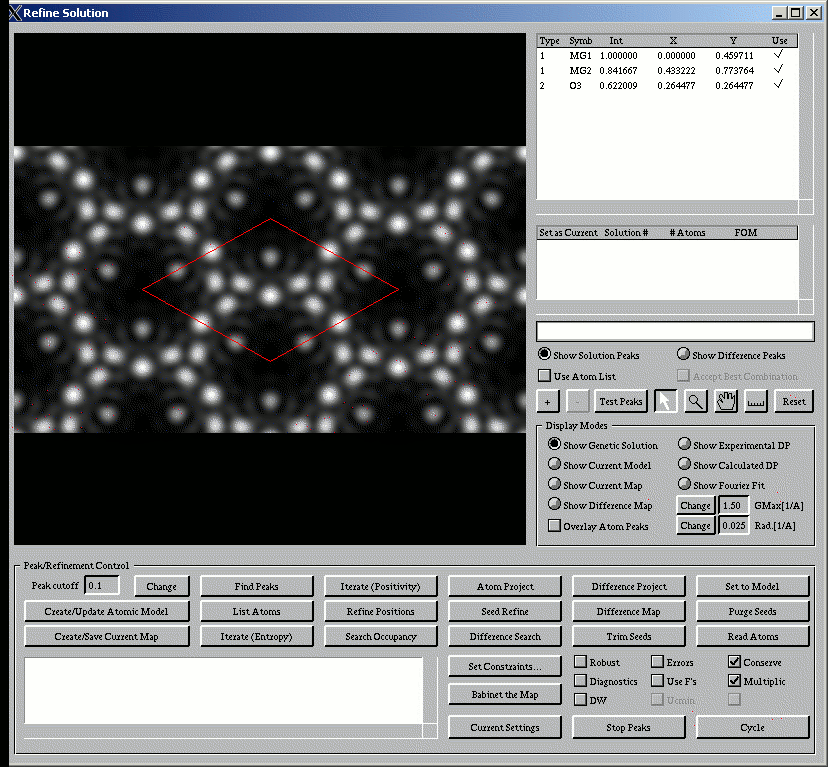
General structure completion methodology:
When you start the program, it will bring up (in the window at the top left) the current direct methods map. An initial peak since will have been done, and an initial guess at the atom locations will appear in the window at the top right. What you do next can, and should vary from case to case; you should probably try different approaches and see if you get the same answer; you might not! In some form, what you will do next is:
- Find Peaks in the initial map
- Attempt to replace these by atoms while simultaneously refining the positions
- Use this information about the atoms to improve the map
- Find atoms that were not included in the previous search, and continue iterating through the process
The main way to find the best combination of atoms is to use Search Occupancy, which will try and find the optimal combination of atomic sites that gives the smallest Figure of Merit (FOM). You will often find the strong scatterers quickly, and these may dominate the direct methods maps. Finding the weaker scatterers can be harder and will depend upon how large are dynamical diffraction effects as well as how good are your measurements.
Following this, a standard method would be to project the map onto the atomic positions that you have found (either the Atom Project or Difference Project commands, followed by a direct methods cycle using either or both Interate (Entropy) and Iterate (Positivity)) then do use Difference Map to find additional possible sites. The Difference Search will test to try and find which of the additional sites improves your FOM. You can also do this manually using the "+" "-" and Test Peaks commands coupled with changing which atom is at which location and adding sites by clicking at different locations. (See the section Atom Testing and Display section below for details about the different commands.)
After this, a reasonable step would be to do a Seed Refinement. This will see which of the various combinations gives the best FOM. Be careful: unless you have bumping constraints set you may get a small FOM that is chemically unreasonable with too many atoms very close to each other.
Tips
1. It is helpful to check the Diagnostics Button and Overlay Atom Peaks. This will allow you to track your progress through the structure refinement. The Overlay Atom Peaks distinguishes between symmetry dependent atoms (outlined in red) and symmetry generated atoms (outlined in blue). At the moment no colors are used to distinguish between different atom types. Although you may select an atom in the map area and the corresponding atom will be highlighted in the atoms list box.
2. The Cycle command iterates through this general procedure (Steps 1-3) and refines the model. While it is rather robust, it is advisable to manual iterate through the refinement procedure.
3. The Current Settings function will list in the display window all of the constraints set on the model, and the specific parameters used in the refinement. Knowing these values is quite helpful when resolving structure.>
4. It is suggested to refine against the R-factor criterion first, since it fits the stronger reflections better than the weaker ones. As the model improves, use the χ2 criterion (check the Errors Button) since it is better at describing the details of the peaks in the fit.
5. The Refine command produces a file Refine.ins which has a format similar to that of a Shelx file and can be read by many standard crystallographic structure display programs. You may want to have a second terminal (or a file manager in windows) open so you can copy this file to one with a different name. You can then (later) read back this solution and others to compare them.
6. It is useful to use the Seed Refinement at various stages. One of the possible solutions may, initially, be rather bad but after moving the atom positions may in fact give a very good fit.
7. Constraints can be very useful. You can vary the strength of them by changing the Weight; if this is small they will be weak, if large they will be strong. You probably want to start of with a small value for these, the default value of 1.0 is OK. Later you might want to increase this.
8. Of the constraints, the most useful in many cases is Bumping which will prevent atoms from coming too close together (in projection). Unfortunately, real structures can have atoms which overlap in projection, so be careful.
9. It is sometimes useful to limit the contents of the atoms in the unit cell (all sites, not the irreducible part of the cell). There are two ways to do this. The first is to use Stoichiometry. The code applies a constraint that the sum of the valences plus the base term in the unit cell is zero. For a bulk material you would use zero for the base value; for a surface 1/2 the charge of the terminating layer. The second method is to use Cell Content which will limit the number of each type of atom between the ranges you input.
Descriptions of the general commands available to the user are listed below and a suggested iteration process (with an example) is provided.
Display Modes
|
Show Genetic Solution |
Shows the map that you started with |
|
Show Current Model |
Displays Current Atom positions with amplitudes and phases |
|
Show Current Map |
Displays Current Atom positions with experimental amplitudes and phases from the model |
|
Show Difference Map |
Displays Difference Map (as defined above) |
|
Show Experimental DP |
Displays experimental diffraction pattern
|
|
Show Calculated DP |
Displays calculated diffraction pattern based on the current model |
|
Show Fourier Fit |
Difference between the Experimental DP and the Calculated DP |
Atoms Testing and Display
|
Atoms Window (Top) |
In this window you have a list of the sites in the current model. In conjunction with the other commands (see below) you can explore manually different assignments of atom types and add/remove atoms. Two particular options:
|
|
Seed Window |
This window contains a list of the different possible solutions that the code has found so far together with their Figure of Merit (FOM). By clicking in the "Set as Current" region you can make any the one currently under consideration |
|
Use Atom List & Test |
If this is checked, and you press on "Test" the current list of atoms will become the default one. If it is unset, then pressing "Test" will report the FOM for the list of atoms. |
|
Show Solution Peaks |
This will show in the top window the atomic positions and types of the current model |
|
Show Difference Peaks |
This will show in the top window the current positions found in a difference map |
|
+ |
By clicking on this button, you activate the cursor so that you can add an atom at a location in the structure shown on the left using the mouse |
|
- |
If you highlight an atom in the list of sites at the top, this can be used to remove an atom |
|
Magnifier/Hand/Ruler Reset |
These allow you to expand the image on the left (Magnifier), move it around (Hand) or measure the distance in Angstroms between to features (Ruler). Reset brings the scale etc back to the original values. |
Refinement Control Commands
|
Create/Update Atomic Model |
This will create an image of the current atomic model (since this may have changed) so it can be viewed. |
|
Create/Save Current Map |
This will create an image of the current map (since this may have changed) so it can be viewed |
|
Find Peaks |
Find peaks and the center-of-mass from current estimate. A list of peaks is kept within the program as the current atomic positions, and also output to a file called "peaks.lst" |
|
List Atoms |
Lists the current atoms (peaks) in the model |
|
Iterate (Entropy) |
Runs another cycle from the current phase in Entropy Mode |
|
Iterate (Positivity) |
Runs another cycle from the current phase in Positivity Mode (map is constrained so that it is positive everywhere, i.e. no negative scattering potential) |
|
Refine Positions |
Refines current atom positions of model. Required to generate the calculated diffraction pattern |
|
Search Occupancy |
Using the current set of peaks, this function searches to find the best approximate Figure of Merit (either an R-factor or χ2 depending on which mode you are refining against) so that matches the experimental data. This produces a list of solutions in the Plausible Solution window. After selecting one of these models, confirm your choice by selecting Create/Update Model |
|
Atom Project |
This function places atoms in locations of peaks in the current model. Given a set of atoms with either variable or fixed heights (all the same), this will replace the peaks in real space by those due to the known atoms, eliminate negative regions and re-correct the moduli in reciprocal space to the experimental values. In effect this projects the solutions onto the set with atoms in certain known positions. A number of cycles of this projection can be applied to improve the current map. |
|
Seed Refine |
Refines all plausible models |
|
Difference Search |
This function looks for a solution which improves the FOM by evaluating the difference peaks in the model and the genetic solution and attempts to place an atom there. It is helpful to use Show Difference Peaks to locate the positions of these features.
|
|
Difference Project |
This is similar to the Atom Project command, but instead looks at the difference between the known atoms and the current map. Since this difference should be atoms, it does an entropy-type projection of this difference then adds back in the known structure. A number of cycles of this projection can be applied to improve the current map |
|
Difference Map |
This command generates the difference map between the current image and current model |
|
Set to Model |
Using the current model (atom positions), this command generates the phases for the current map using the experimental amplitudes and the calculated phases. |
|
Trim Seeds |
Removes the remaining solutions (besides the current model) from the plausible model repository. New difference searches are the isolated to just that current model. |
|
Purge Seeds |
A more forceful Trim Seeds command. |
|
Read Atoms |
Can read in a Case.ins file with atom positions |
Refinement Options
|
Robust |
Refines on the moduli, as against the squares. If, for instance, you want to do an R1 refinement, use this flag. If you have good errors, and at the start of the analysis you probably do not want to use this. |
|
Errors |
When you check this box, the code will read in a file called "full.hkl" which has a general format "h k l Value Error" where both Value and Error can be either intensities or structure factors. Note: if they are structure factors then you must use the "Use F's" flag. Subsequent refinements will use these values, with the errors if this box is checked, without them if it is not. Note: the values in full.hkl can (often are) be much larger than the values used in the direct methods analysis. A Chi**2 analysis would be done with errors and "Robust" unchecked; a Chi analysis with both checked. |
|
Conserve |
This will apply a constraint that the experimental and calculated intensities are similar in absolute terms when you are doing a Chi**2 or Chi analysis. Without it, there is a danger that only the weak reflections are being fit and the number is misleadingly small. |
|
Use F's |
This will refine on the structure factors, as against intensities |
|
Multiplicity |
This will refine multiplying each reflection by the number of equivalent (hkl) reflections. In effect, this refines on the total diffracted intensity. For an R1 or R2 refinement this is probably a good idea; it is probably a bad idea for a Chi**2 or Chi refinement. |
|
DW |
When checked, the temperature factors will be partially refined as isotropic variables for each atom type. |
|
Diagnostics |
This will provide additional information. |